Non Linear Approximation
- Introduction
- Non Linear Least Square Approximation
- Levenberg-Marquardt algorithm
- Uniform Approximation - the Remez Algorithm
Introduction
We are likely all familiar with the concept of curve-fitting — sometimes, we, as humans,
can recognize certain patterns in data about which the data itself is agnostic.
If we look at a chart like the following:
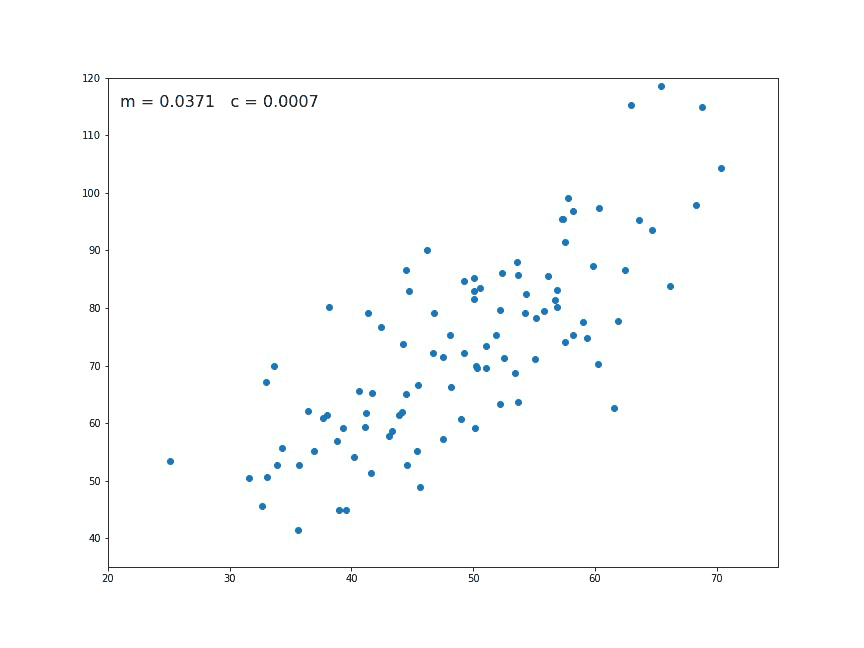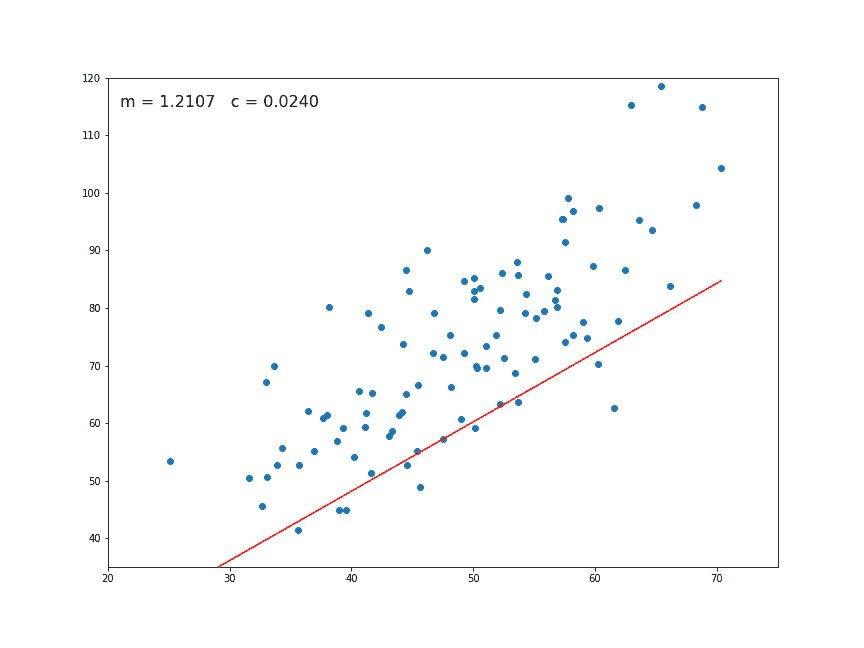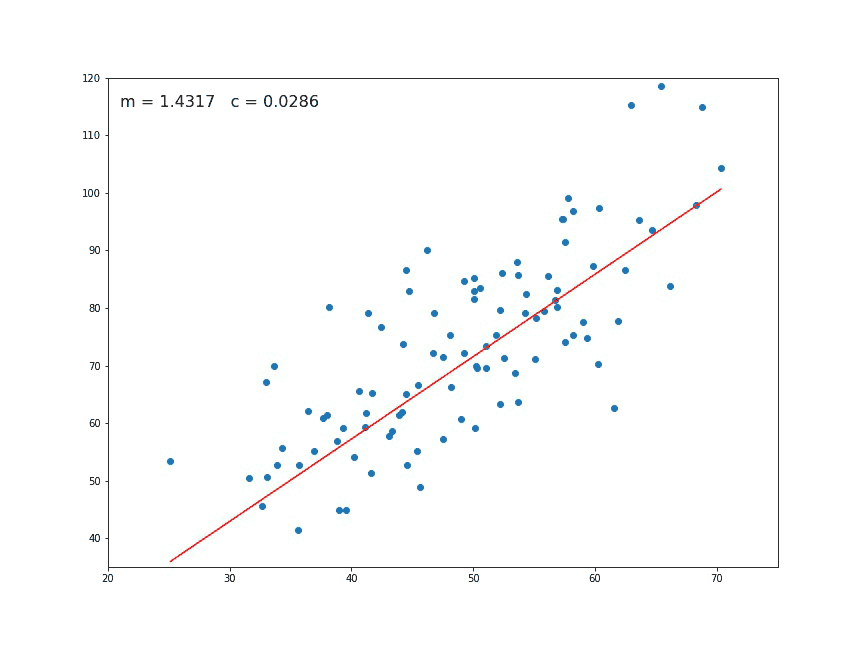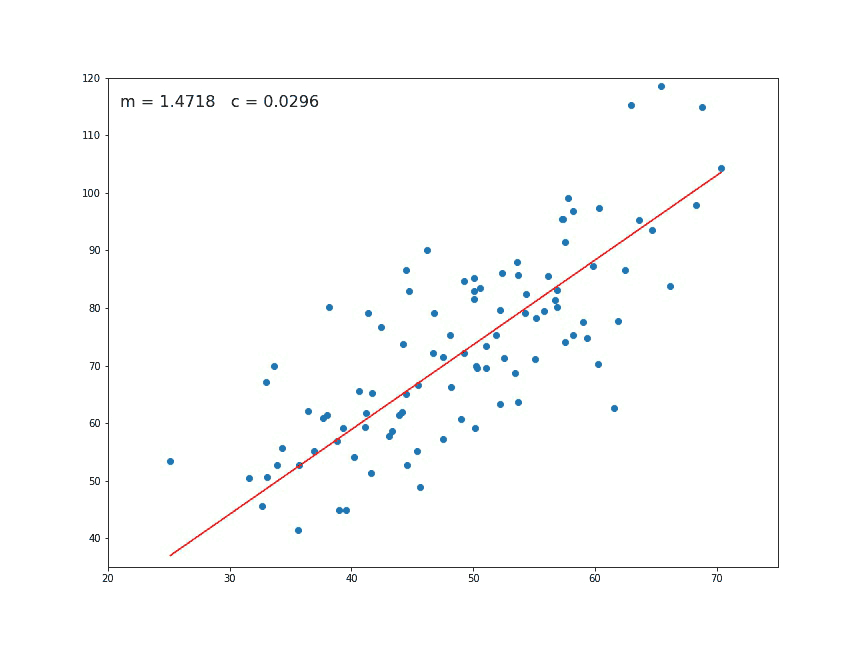 We think immediately of a linear relationship. But what tools can we use to precisely determine a line from these individual points of data?
Intuitively speaking, we can all figure this out pretty easily — we want to find the line that is minimally far from every single one of these points.
The way this is actually implemented is through a method known as least squares regression.
This form of regression is very powerful, and is widely used in applications from signals processing to economics research.
Least squares fitting finds the best curve to fit a set of points through
minimizing the sum of the squares of the offsets of each point from the curve.
We think immediately of a linear relationship. But what tools can we use to precisely determine a line from these individual points of data?
Intuitively speaking, we can all figure this out pretty easily — we want to find the line that is minimally far from every single one of these points.
The way this is actually implemented is through a method known as least squares regression.
This form of regression is very powerful, and is widely used in applications from signals processing to economics research.
Least squares fitting finds the best curve to fit a set of points through
minimizing the sum of the squares of the offsets of each point from the curve.
Non Linear Least Square Approximation
The goal of nonlinear least squares (LS) regresion is to
model the nonlinear relationship between the dependent variable $b$ and independent variables in
${\bf x}=[x_1,\cdots,x_d]^T$,
based on the given training dataset
${\cal P}=\{({\bf x}_j, y_j),\;j=1,\cdots,p\}$,
by a nonlinear regression function parameterized
by a set of $M$ parameters represented by
${\bf\theta}=[\theta_1,\cdots,\theta_M]^T$
\begin{equation}
\hat{y}=f(x_1,\cdots,x_d,\;\theta_1,\cdots,\theta_M)
=f({\bf x},{\bf\theta})
\end{equation}
While the specific form of the nonlinear function is typically
determined based on certain prior knowledge, we need to find
the parameeter ${\bf\theta}$ for the model to fit the
training dataset ${\cal P}$, so that, its output $\hat{\bf x}$
matches the ground truth labeling ${\bf b}$ in the training set,
i.e., the residual ${\bf r}={\bf y}-\hat{\bf y}$
${\bf r}={\bf y}-\hat{\bf y}$ is ideally zero:
\begin{equation}
{\bf r}({\bf\theta})
=\left[\begin{array}{c}r_1\\\vdots\\r_N\end{array}\right]
=\left[\begin{array}{c}y_1\\\vdots\\y_N\end{array}\right]
-\left[\begin{array}{c}f_1({\bf\theta})\\
\vdots\\f_N({\bf\theta}) \end{array}\right]
={\bf y}-\hat{\bf y}({\bf\theta})
={\bf y}-{\bf f}({\bf X},{\bf\theta})={\bf0}
\end{equation}
where
$\hat{y}({\bf x}_n,{\bf\theta})=f_n({\bf\theta})=f({\bf x}_n,{\bf\theta})$
is the predicted output by the regression function based on the
j-th data point ${\bf x}_j$. We see that the regression problem
is equivalent to solving a nonlinear equation system of $p$
equations for the $M$ unknown parameters in ${\bf\theta}$. As
typically there are many more data samples than the unknown
parameters, i.e., $p\gg M$, the equation ${\bf r}={\bf0}$
is over-overconstrained without an exact solution. We therefore
consider the the optimal parameter ${\bf\theta}^*$ that minimizes
the sum-of-squared error based on the residual ${\bf r}$:
\begin{equation}
\varepsilon({\bf\theta})=\frac{1}{2}||{\bf r}||^2
=\frac{1}{2}\sum_{n=1}^N r_n^2
=\frac{1}{2}\sum_{n=1}^N[y_n-f_n({\bf\theta})]^2
\end{equation}
To do so, we first consider the gradient descent method based
on the gradient of this error function:
\begin{eqnarray}
{\bf g}_\varepsilon({\bf\theta})
&=&\frac{d}{d{\bf\theta}}\varepsilon({\bf\theta})
=\frac{d}{d{\bf\theta}}\left( \frac{1}{2}\sum_{n=1}^N r_n^2\right)
=\sum_{n=1}^N \frac{d\,r_n}{d{\bf\theta}}\;r_n
\nonumber\\
&=&-\sum_{n=1}^N \frac{d}{d{\bf\theta}} f_n({\bf\theta})
[y_n-f_n({\bf\theta})] =-{\bf J}_f^T\;{\bf r}
\end{eqnarray}
where ${\bf J}_f$ is the $N\times M$ Jacobian matrix ${\bf J}_f$ of ${\bf f}({\bf X},{\bf\theta})$.
The mth component of the gradient vector is
\begin{eqnarray}
g_m({\bf\theta})&=&\frac{\partial}{\partial\theta_m}\varepsilon({\bf\theta})
=\frac{1}{2} \frac{\partial}{\partial\theta_m}||{\bf r}||^2
=\frac{1}{2} \sum_{n=1}^N \frac{\partial}{\partial\theta_m}r_n^2
=\sum_{n=1}^N \frac{\partial r_n}{\partial\theta_m}\;r_n
\nonumber\\
&=&\sum_{n=1}^N
\frac{\partial f_n({\bf\theta})}{\partial\theta_m}
[f_n({\bf\theta})-y_n]
=\sum_{n=1}^N J_{nm}\,[f_n({\bf\theta})-y_n]
\sum_{n=1}^N J_{nm}\,r_n
\;\;\;\;\;\;\;\;(m=1,\cdots,M)
\end{eqnarray}
where $J_{nm}=\partial\,f_n({\bf\theta})/\partial\theta_m$ is
the element in the nth row and mth column of ${\bf J}_f$,
which is simply the negative version of Jacobian of
${\bf r}={\bf y}-{\bf f}({\bf X},{\bf\theta})$
\begin{equation}
{\bf J}_r({\bf\theta})=\frac{d}{d{\bf\theta}}{\bf r}({\bf\theta})
=\frac{d}{d{\bf\theta}} [ {\bf y}-{\bf f}({\bf X},{\bf\theta}) ]
=-\frac{d}{d{\bf\theta}} {\bf f}({\bf X},{\bf\theta})
=-{\bf J}_f({\bf\theta})
\end{equation}
The optimal parameters in ${\bf\theta}$
that minimizes $\varepsilon({\bf\theta})$
can now be found iteratively by the gradient descent method:
\begin{equation}
{\bf\theta}_{n+1}={\bf\theta}_n-\delta_n {\bf g}_\varepsilon({\bf\theta}_n)
={\bf\theta}_n-\delta_n {\bf J}_f^T\;{\bf r}
\end{equation}
Setting the value for the step size $\delta_n$ properly may
be tricky, as it depends on the specific landscape of the
error hypersurface $\varepsilon({\bf\theta})$. If $\delta_n$ is too large,
a minimum may be skipped over, but if $\delta_n$ is too small,
the convergence of the iteration may be too slow.
This gradient method is also called batch gradient descent (BGD), as the gradient is calculated based on the entire batch of all $p$ data points in the dataset ${\cal P}$ in each iteration. When the size of the dataset is large, BGD may be slow due to the high computational complexity. In this case the method of stochastic gradient descent (SGD) can be used, which approximates the gradient based on only one of the $p$ data points uniformly randomly chosen from the dataset in each iteration, i.e., the summation contains only one term corresponding to the chosen sample ${\bf x}_j$: \begin{equation} {\bf g}_n({\bf\theta})=\frac{d}{d{\bf\theta}} \frac{1}{2} r_n^2 =r_n\frac{d\,r_n}{d{\bf\theta}} \end{equation} The expectation of ${\bf g}_n$ based on one data point is the same as the true gradient (scaled by $1/p$) based on all $p$ data points in the dataset \begin{equation} E[ {\bf g}_n({\bf\theta}) ] =\sum_{n=1}^N P({\bf x}_n) {\bf g}_n({\bf\theta}) =\sum_{n=1}^N \frac{1}{N}{\bf g}_n({\bf\theta}) =\frac{1}{N}\sum_{n=1}^N \frac{dr_n}{d{\bf\theta}}\;r_n =\frac{1}{N}{\bf g}({\bf\theta}) \end{equation}
As the data are typically noisy, the SGD gradient based on only one data point may be in a direction very different from the true gradient based on the mean of all $p$ data points, and the resulting search path in the M-D space may be in a very jagged zig-zag shape, and the iteration may converge slowly. However, as the computational complexity in each iteration is much reduced, we can afford to carry out a large number of iterations with an overall search path generally coinciding with that of the BGD method.
Some trade-off between BGD and SGD can be made so that the gradient is estimated based on more than one but less than all $p$ data points in the dataset, the resulting method, called mini-batch gradient descent, can take advantage of both methods. The methods of stochastic and mini-batch gradient descent are widely used in machine learning, such as in the method of artificial neural networks, which can be treated as a nonlinear regression problem.
Alternatively, the optimal parameter ${\bf\theta}^*$ of the nonlinear regression function $f({\bf x},{\bf\theta})$ can also be found by the Gauss-Newton method, which directly solves the nonlinear over-determined equation system ${\bf r}({\bf\theta})={\bf y}-{\bf f}({\bf X},{\bf\theta})={\bf0}$, based on the iterative Newton-Raphson method. \begin{equation} {\bf\theta}_{n+1} ={\bf\theta}_n-{\bf J}_r^-\;{\bf r} ={\bf\theta}_n-({\bf J}_r^T{\bf J}_r)^{-1}{\bf J}_r^T\;{\bf r} ={\bf\theta}_n+({\bf J}_f^T{\bf J}_f)^{-1}{\bf J}_f^T\;{\bf r} \end{equation}
The last equality is due to the fact that ${\bf J}_r=-{\bf J}_f$. Here ${\bf J}_f^T{\bf J}_f$ in the pseudo-inverse is actually an approximation of the Hessian ${\bf H}_\varepsilon$ of the error function $\varepsilon({\bf\theta})$, and $-{\bf J}^T_f{\bf r}={\bf g}_\varepsilon({\bf\theta})$ is the gradient of the error function. So the iteration above is essentially: \begin{equation} {\bf\theta}_{n+1} ={\bf\theta}_n-{\bf J}_r^-\;{\bf r} \approx {\bf\theta}_n -{\bf H}_\varepsilon^{-1}({\bf\theta}_n){\bf g}_\varepsilon({\bf\theta}_n) \end{equation} same as the Newton-Raphson method based on the second order Hessian as well as the first oder gradient. We therefore see that the Gauss-Newton method is actually for minimizing the sum-of-squares error is essentially the same as the Newton-Raphson method, with the only difference that the Hessian ${\bf H}_\varepsilon \approx {\bf J}_f^T{\bf J}_f$ as the second order derivative of the error function $\varepsilon({\bf\theta})$ is approximated by the Jacobian ${\bf J}_f{\bf\theta}$ as the first order derivative of the regression function ${\bf f}({\bf x},{\bf\theta})$, making the implementation computationally easier.
In the special case of linear regression function ${\bf f}({\bf w})={\bf x}^T{\bf w}$, we have: \begin{equation} \hat{\bf y}={\bf f}({\bf X},{\bf w}) =\left[\begin{array}{c}f({\bf x}_1,{\bf w})\\ \vdots\\f({\bf x}_N,{\bf w})\end{array}\right] =\left[\begin{array}{c}{\bf x}_1^T\\\vdots\\ {\bf x}_N^T\end{array}\right]{\bf w} =[{\bf x}_1,\cdots,{\bf x}_N]^T{\bf w}={\bf X}^T{\bf w} \end{equation} where ${\bf w}=[w_0,\,w_1,\cdots,w_d]^T$ and ${\bf x}_i=[1, x_{i1},\cdots,x_{id}]^T$ are both augmented $d+1$ dimensional vectors and ${\bf X}=[{\bf x}_1,\cdots,{\bf x}_N]$. The Taylor series is \begin{equation} f({\bf X},{\bf w}+\Delta{\bf w})={\bf X}^T({\bf w}+\Delta{\bf w}) ={\bf X}^T{\bf w}+{\bf X}^T\Delta{\bf w} \end{equation} and the Jacobian is \begin{equation} {\bf J}_f({\bf w})={\bf X}^T \end{equation} Now Eq. () becomes \begin{eqnarray} {\bf w}_{n+1}&=&{\bf w}_n+({\bf X}^T)^-({\bf y}-{\bf X}^T{\bf w}_n) ={\bf w}_n+({\bf X}{\bf X}^T)^{-1}{\bf X}({\bf y}-{\bf X}^T{\bf w}_n) \nonumber\\ &=&{\bf w}_n+({\bf X}{\bf X}^T)^{-1}{\bf X}{\bf y} -({\bf X}{\bf X}^T)^{-1}({\bf X}{\bf X}^T){\bf w}_n \nonumber\\ &=&({\bf X}{\bf X}^T)^{-1}{\bf X}{\bf y}=({\bf X}^T)^-{\bf y} \end{eqnarray} i.e., the optimal parameter ${\bf w}^*$ can be found in a single step without iteration, same as that in Eq. ().
The Python code for the essential parts of the algorithm is
listed below, where data(:,1) and data(:,2) are the
observed data containing $p$ samples $x_1,\cdots,x_p$
in the first column and their corresponding function values $y_1,\cdots,y_p$
in the second column. In the code segment below, the regression
function (e.g., an exponential model) is symbolically defined, and
then a function NLregression is called to estimate the
function parameters:
sym x; % symbolic variables
a=sym('a',[3 1]); % symbolic parameters
f=@(x,a)a(1)*exp(a(2)*x)+a(3)| % symbolic function
[a er]=NLregression(data(:,1),data(:,2),f,3); % call regression
The code below estimates the $M$ parameters of the regression function based on Newton's method.
function [a er]=NLregression(x,y,f,M) % nonlinear regression
N=length(x); % number of data samples
a=sym('a',[M 1]); % M symbolic parameters
F=[];
for i=1:N
F=[F; f(x(i),a)]; % evaluate function at N given data points
end
J=jacobian(F,a); % symbolic Jacobian
J=matlabFunction(J); % convert to a true function
F=matlabFunction(F);
a1=ones(M,1); % initial guesses for M parameters
n=0; % iteration index
da=1;
while da>tol % terminate if no progress made
n=n+1;
a0=a1; % update parameters
J0=J(a0(1),a0(2)); % evaluate Jacobian based on a0
Jinv=pinv(J0); % pseudo inverse of Jacobian
r=y-F(a0(1),a0(2)); % residual
a1=a0+Jinv*r; % Newton iteration
er=norm(r); % residual error
da=norm(a0-a1); % progress
end
end
where tol is a small value for tollerance, such as $10^{-6}$.
Example 1
Consider three sets of data (shown as the solid dots in the three columns in the plots below) are fitted by three different models (shown as the open circles in the plots):
- Linear model: $y=f_1(x)=ax+b$
- quadratic model: $y=f_2(x)=ax^2+bx+c$
- Exponential model: $y=f_3(x)=a\,e^{bx}+c$
These are the parameters for the three models to fit each of the three datasets: \begin{equation} \begin{array}{r||r||r|r|r||r}\hline & Parameters: & a & b & c & error \\\hline \hline & Model\;1: & 30.337 & 51.652 & & 7.4294 \\ Dataset\;1 & Model\;2: & 0.076 & 30.034 & 51.580 & 7.4223 \\ & Model\;3: &5933.214 & 0.005 & -5881.634 & 7.4221 \\\hline & Model\;1: & 24.519 & 20.089 & & 21.0362 \\ Dataset\;2 & Model\;2: & 4.459 & 6.625 & 15.855 & 8.4559 \\ & Model\;3: & 29.791 & 0.346 & -10.202 & 9.7402 \\\hline & Model\;1: & 32.608 & 22.008 & & 53.0724 \\ Dataset\;3 & Model\;2: & 10.391 & -9.089 & 12.140 & 28.3211 \\ & Model\;3: & 2.197 & 0.885 & 27.241 & 8.3393 \\\hline \end{array} \nonumber \end{equation}
We see that the first dataset can be fitted by all three models equally well with similar error (when $a\approx 0$ in model 2, it is approximately a linear functions), the second dataset can be well fitted by either the second or the third model, while the third dataset can only be fitted by the third exponential model.
Example 2
Based on the following three functions with four parameters $a=1,\;b=2,\;c=3,\;d=4$, three sets of 2-D data points are generated (with some random noise added). \begin{eqnarray} f_1({\bf x})&=&ax_1+bx_2-c \nonumber\\ f_2({\bf x})&=&\exp(ax_1+bx_2)-cx_1x_2+d \nonumber\\ f_3({\bf x})&=&ax_1^2-bx_1x_2+cx_2^2+d \end{eqnarray}
Then each of three datasets are fitted by the three functions as models while the optimal parameters are found by the Gauss-Newton method. \begin{equation} \begin{array}{r||r||r|r|r|r||r}\hline & Parameters: & a & b & c & d & error \\\hline \hline Dataset\;1 & Model\;1: & 0.98 & 1.97 & 3.00 & & 4.13 \\ & Model\;2: & 0.70 & 1.30 & 1.21 & -4.42 & 6.87 \\ & Model\;3: & -0.19 & 0.73 & -0.12 & -3.01 & 18.03 \\\hline Dataset\;2 & Model\;1: & 1.84 & 3.40 & -6.09 & & 14.70 \\ & Model\;2: & 0.99 & 2.01 & 2.97 & 3.98 & 3.75 \\ & Model\;3: & 0.37 & 1.13 & 2.66 & 4.87 & 32.45 \\\hline Dataset\;3 & Model\;1: & -0.19 & 0.30 & -5.34 & & 16.82 \\ & Model\;2: & -0.29 & 0.23 & 2.00 & 4.30 & 13.51 \\ & Model\;3: & 0.94 & 2.05 & 2.99 & 4.02 & 4.09 \\\hline \end{array} \nonumber \end{equation}
Levenberg-Marquardt algorithm
The Levenberg-Marquardt algorithm (aka damped least-squares method) can be considered as trade-off between the Gauss-Newton method and the gradient descent method. While all such methods can be used to minimize an objective function, such as the LS error $\varepsilon({\bf a})$ of a data fitting (modeling) problem, the Levenberg-Marquardt method is more robust. Even if the initial guess is far from the solution corresponding to the minimum of the objective function, the iteration can still converge toward the solution.
Specifically, the normal equation obtained in the Gauss-Newton method \begin{equation} \left({\bf J}^T{\bf J}\right)\,\triangle{\bf a}={\bf J}^T({\bf y}-{\bf f}({\bf a})) \end{equation} is modified by adding an extra term proportional to the identity matrix ${\bf I}$: \begin{equation} \left({\bf J}^T{\bf J}+\lambda {\bf I}\right)\,\triangle{\bf a} ={\bf J}^T ({\bf y}-{\bf f}({\bf a})) \end{equation} Here $\lambda$ is the non-negative damping factor, which is to be adjusted at each iteration. Solving for $\triangle{\bf a}$, we get \begin{equation} \triangle{\bf a}=({\bf J}^T{\bf J}+\lambda{\bf I})^{-1} \left[{\bf J}^T ({\bf y}-{\bf f}({\bf a}))\right] \end{equation} When $\lambda$ is close to zero, the above is essentially the Gauss-Newton method discussed previously. But when $\lambda$ is large, ${\bf J}^T{\bf J}$ becomes insignificant, $\triangle{\bf a}$ is approximately proportional to ${\bf J}^T({\bf y}-{\bf f}({\bf a}))$, which is the negative gradient of $\varepsilon({\bf a})$ \begin{equation} {\bf g}_\varepsilon({\bf a}) =-2\,{\bf J}^T({\bf y}-{\bf f}({\bf a})) \end{equation} We realize that the iteration is actually a combination of Gauss-Newton method and gradient descent method. The parameter $\lambda$" is used to emphasize either of the two methods for the same purpose of minimizing $\varepsilon({\bf a})$. Initially $\lambda$ can be set to a small value so that the iteration is dominated by the Gauss-Newton method. But if the objective function is reduced too slowly, the value of $\lambda$ is increased, the iteration is then switched to the gradient descent method.
References
- C. Eckart, G. Young, The approximation of one matrix by another of lower rank. Psychometrika, Volume 1, 1936, Pages 211–8. doi:10.1007/BF02288367
- C. de Boor (1978) A Practical Guide to Splines. Springer-Verlag.
- G.D. Knott (2000) Interpolating Cubic Splines. Birkhäuser.
- H.J. Nussbaumer (1981) Fast Fourier Transform and Convolution Algorithms. Springer-Verlag.
- H. Späth (1995) One Dimensional Spline Interpolation. AK Peters.